Difference between revisions of "Project Week 25/Breast Segmentation in DCE-MRI via Deep Learning Approaches"
| Line 6: | Line 6: | ||
==Key Investigators== | ==Key Investigators== | ||
<!-- Key Investigator bullet points --> | <!-- Key Investigator bullet points --> | ||
| − | *[https://www.researchgate.net/profile/Gabriele_Piantadosi Gabriele Piantadosi] (University Federico II di Napoli, Italy) | + | *[https://www.researchgate.net/profile/Gabriele_Piantadosi Gabriele Piantadosi] (gabriele.piantadosi@unina.it - University Federico II di Napoli, Italy) |
==Background== | ==Background== | ||
In recent years Dynamic Contrast Enhanced-Magnetic Resonance Imaging (DCE-MRI) has gained popularity as an important complementary diagnostic methodology for early detection of breast cancer. It has demonstrated a great potential in the screening of high-risk women, in staging newly diagnosed breast cancer patients and in assessing therapy effects thanks to its minimal invasiveness and to the possibility to visualise 3D high resolution dynamic (functional) information not available with conventional RX imaging. | In recent years Dynamic Contrast Enhanced-Magnetic Resonance Imaging (DCE-MRI) has gained popularity as an important complementary diagnostic methodology for early detection of breast cancer. It has demonstrated a great potential in the screening of high-risk women, in staging newly diagnosed breast cancer patients and in assessing therapy effects thanks to its minimal invasiveness and to the possibility to visualise 3D high resolution dynamic (functional) information not available with conventional RX imaging. | ||
| − | Among the major issues in developing CAD systems for breast DCE-MRI there are: (a) the detection of | + | Among the major issues in developing CAD systems for breast DCE-MRI there are: (a) the detection of suspicious regions of interest (ROIs) as sensibly as possible, while simultaneously minimising the number of false alarms [1] and (b) the classification of each segmented ROI according to its aggressiveness. This task is made harder by the peculiarity of DCE-MRI breast examinations: breast movements due to inspiration, a huge diversity of lesion types. |
==Project Description== | ==Project Description== | ||
| − | + | Breast Cancer analysis could be approached as a classification problem via Computer Aided Detection (CAD) system. | |
| − | + | [[File:Typical Computer Aided Detection System.png|thumb|center|upright=2.0|Typical Computer Aided Detection System]] | |
| − | [[File: | + | '''Breats-Mask Extraction''' |
| + | |||
| + | With the aim of reducing the computational cost of further steps and attenuate noise caused by extraneous voxel (VOlumetric piXEL) a binary mask representing only breast parenchyma and excluding background and other tissues is extracted. Segmentation of breast parenchyma has been addressed relying on fuzzy binary clustering, breast anatomical priors and morphological refinements. | ||
| + | |||
| + | '''My Previous Proposal (ICPR2016 [1])''' | ||
| + | |||
| + | The segmentation of breast parenchyma has been addressed relying on fuzzy binary clustering, breast anatomical priors and morphological refinements. In breast segmentation, the most difficult issue to address is discriminating the breast parenchyma from the pectoral muscle since signal intensities, textures and anatomical structures of these tissues are very close each other. | ||
| + | |||
| + | [[File:A Breast-Mask extracted from DCE-MRI data.png|thumb|center|upright=2.0|A Breast-Mask extracted from DCE-MRI data]] | ||
| + | |||
| + | The proposed breast mask extraction approach overcomes the issue by mixing geometrical-based and pixel-based approaches. It relies on geometrical anatomical priors to take advance of anatomical knowledge of the breast key points and uses a pixel base segmentation to obtain the best threshold for each border. | ||
| + | |||
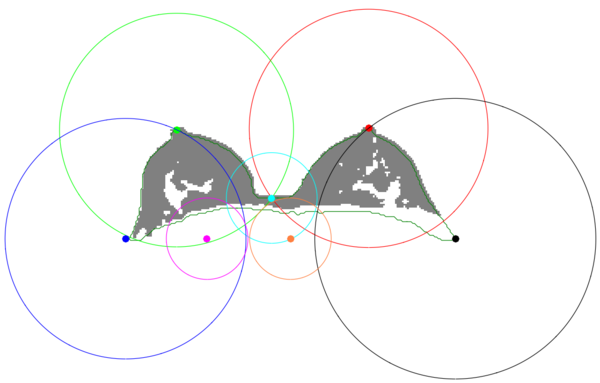
| + | [[File:Anatomical key-points.png|thumb|center|upright=2.0|Anatomical key-points]] | ||
| + | |||
| + | It uses a pixel-based Fuzzy C-Means (FCM) clustering to shift the breast mask extraction from a simple grey-level based segmentation to a membership probability one. Moreover, it exploits novel geometrical consideration to weight the classes membership probability according to the breast anatomy. | ||
| + | |||
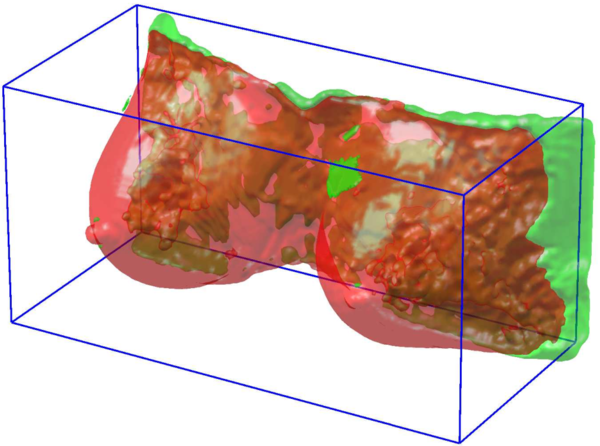
| + | [[File:Result of FCM segmentation (in red) overlapped to the Ground Truth (green).png|thumb|center|upright=2.0|Result of FCM segmentation (in red) overlapped to the Ground Truth (green)]] | ||
| + | |||
| + | The result is an automated procedure, able to extract an accurate breast mask without any prior information on the patient dataset (as in the case of atlases). | ||
| + | |||
| + | '''Deep Proposal''' | ||
| + | |||
| + | The novel proposed lesion detection module performs the breast segmentation relying on deep approaches such as those proposed in [4, 5]. In particular, “u-net” provides segmentation via deep Convolutional Neural Networks (dCNN). | ||
| + | |||
| + | [[File:U-Net architecture for image segmentation.png|thumb|center|upright=3.0|U-Net architecture for image segmentation]] | ||
| + | |||
| + | Segmentation has been performed in Axial projection | ||
| − | |||
| − | |||
{| class="wikitable" | {| class="wikitable" | ||
| Line 30: | Line 54: | ||
|- style="vertical-align:top;" | |- style="vertical-align:top;" | ||
|<!-- Objective bullet points --> | |<!-- Objective bullet points --> | ||
| − | * | + | *Breast segmentation of 4D DCE-MRI volumes via deep approaches [1]. |
| − | [ | ||
*Results evaluation with 42 patients provided of manually segmented ground-truth. | *Results evaluation with 42 patients provided of manually segmented ground-truth. | ||
| Line 42: | Line 65: | ||
|<!-- Progress and Next steps (fill out at the end of project week), bullet points --> | |<!-- Progress and Next steps (fill out at the end of project week), bullet points --> | ||
| − | Relying on the segmented regions of interest (ROIs) a lesion malignity assessment via deep approaches should be performed. | + | *Include the model in a Slicer Module. |
| + | *Relying on the Breast Mask Lesion detection via Deep approaches should be performed [2]. | ||
| + | *Relying on the segmented regions of interest (ROIs) a lesion malignity assessment via deep approaches should be performed [3]. | ||
|} | |} | ||
| + | |||
| + | |||
| + | ==Preliminary Results== | ||
| + | {| class="wikitable" | ||
| + | ! style="font-weight: bold;" | Approach | ||
| + | ! style="font-weight: bold;" | Validation | ||
| + | ! style="font-weight: bold;" | Dice | ||
| + | |- | ||
| + | | fuzzy c-means | ||
| + | | leave one patient out | ||
| + | | 83.25 | ||
| + | |- | ||
| + | | fuzzy c-means + anatomical priors | ||
| + | | leave one patient out | ||
| + | | 88.85 | ||
| + | |- | ||
| + | | fuzzy c-means + anatomical priors + postprocessing | ||
| + | | leave one patient out | ||
| + | | 91.36 | ||
| + | |- | ||
| + | | "u-net" without batch-normalisation | ||
| + | | 30 epochs (hold-out) | ||
| + | | 76.58 | ||
| + | |- | ||
| + | | "u-net" with batch-normalisation | ||
| + | | 30 epochs (hold-out) | ||
| + | | 82.19 | ||
| + | |} | ||
==Background and References== | ==Background and References== | ||
| − | + | 1. Marrone, S., Piantadosi, G., Fusco, R., Petrillo, A., Sansone, M., & Sansone, C. (2016, December). Breast Segmentation using Fuzzy C-Means and anatomical priors in DCE-MRI. In Pattern Recognition (ICPR), 2016 23rd International Conference on (pp. 1472-1477). IEEE. | |
| + | |||
| + | 2. Marrone, S., Piantadosi, G., Fusco, R., Petrillo, A., Sansone, M., & Sansone, C. (2013, September). Automatic lesion detection in breast DCE-MRI. In International Conference on Image Analysis and Processing (pp. 359-368). Springer Berlin Heidelberg. | ||
| + | |||
| + | 3. Piantadosi, G., Fusco, R., Petrillo, A., Sansone, M., & Sansone, C. (2015). LBP-TOP for volume lesion classification in breast DCE-MRI. In 18th International Conference on Image Analysis and Processing, ICIAP 2015. Springer Verlag. | ||
| + | |||
| + | 4. Long, J., Shelhamer, E., & Darrell, T. (2015). Fully convolutional networks for semantic segmentation. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (pp. 3431-3440). | ||
| + | |||
| + | 5. Ronneberger, O., Fischer, P., & Brox, T. (2015). U-net: Convolutional networks for biomedical image segmentation. arXiv preprint arXiv:1505.04597. | ||
Revision as of 12:04, 30 June 2017
Home < Project Week 25 < Breast Segmentation in DCE-MRI via Deep Learning ApproachesBack to Projects List
Breast Analysis in DCE-MRI
Key Investigators
- Gabriele Piantadosi (gabriele.piantadosi@unina.it - University Federico II di Napoli, Italy)
Background
In recent years Dynamic Contrast Enhanced-Magnetic Resonance Imaging (DCE-MRI) has gained popularity as an important complementary diagnostic methodology for early detection of breast cancer. It has demonstrated a great potential in the screening of high-risk women, in staging newly diagnosed breast cancer patients and in assessing therapy effects thanks to its minimal invasiveness and to the possibility to visualise 3D high resolution dynamic (functional) information not available with conventional RX imaging. Among the major issues in developing CAD systems for breast DCE-MRI there are: (a) the detection of suspicious regions of interest (ROIs) as sensibly as possible, while simultaneously minimising the number of false alarms [1] and (b) the classification of each segmented ROI according to its aggressiveness. This task is made harder by the peculiarity of DCE-MRI breast examinations: breast movements due to inspiration, a huge diversity of lesion types.
Project Description
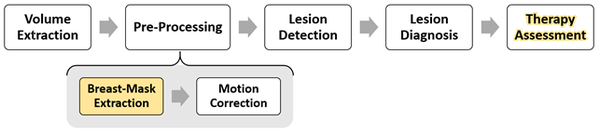
Breast Cancer analysis could be approached as a classification problem via Computer Aided Detection (CAD) system.
Breats-Mask Extraction
With the aim of reducing the computational cost of further steps and attenuate noise caused by extraneous voxel (VOlumetric piXEL) a binary mask representing only breast parenchyma and excluding background and other tissues is extracted. Segmentation of breast parenchyma has been addressed relying on fuzzy binary clustering, breast anatomical priors and morphological refinements.
My Previous Proposal (ICPR2016 [1])
The segmentation of breast parenchyma has been addressed relying on fuzzy binary clustering, breast anatomical priors and morphological refinements. In breast segmentation, the most difficult issue to address is discriminating the breast parenchyma from the pectoral muscle since signal intensities, textures and anatomical structures of these tissues are very close each other.
The proposed breast mask extraction approach overcomes the issue by mixing geometrical-based and pixel-based approaches. It relies on geometrical anatomical priors to take advance of anatomical knowledge of the breast key points and uses a pixel base segmentation to obtain the best threshold for each border.
It uses a pixel-based Fuzzy C-Means (FCM) clustering to shift the breast mask extraction from a simple grey-level based segmentation to a membership probability one. Moreover, it exploits novel geometrical consideration to weight the classes membership probability according to the breast anatomy.
The result is an automated procedure, able to extract an accurate breast mask without any prior information on the patient dataset (as in the case of atlases).
Deep Proposal
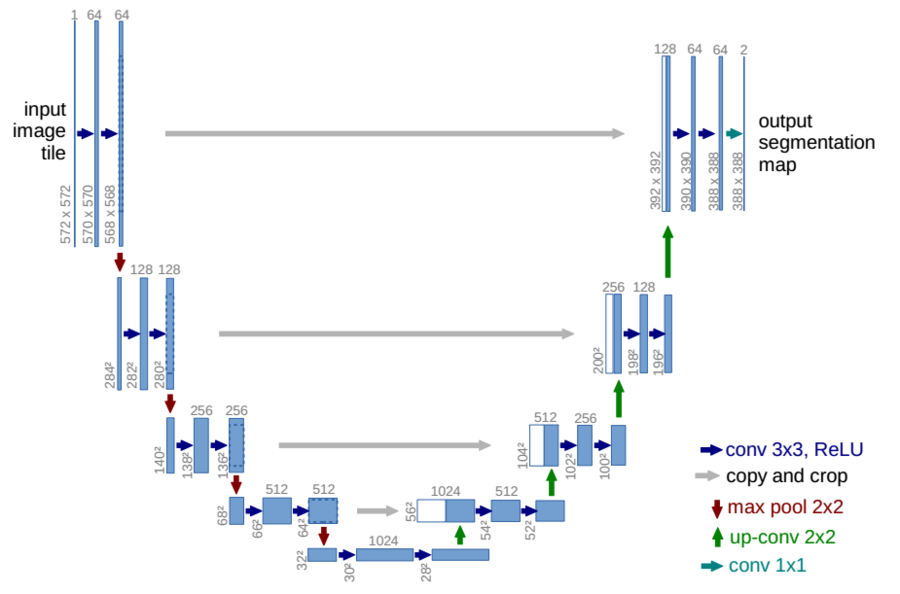
The novel proposed lesion detection module performs the breast segmentation relying on deep approaches such as those proposed in [4, 5]. In particular, “u-net” provides segmentation via deep Convolutional Neural Networks (dCNN).
Segmentation has been performed in Axial projection
| Objective | Approach and Plan | Progress and Next Steps |
|---|---|---|
|
|
|
Preliminary Results
| Approach | Validation | Dice |
|---|---|---|
| fuzzy c-means | leave one patient out | 83.25 |
| fuzzy c-means + anatomical priors | leave one patient out | 88.85 |
| fuzzy c-means + anatomical priors + postprocessing | leave one patient out | 91.36 |
| "u-net" without batch-normalisation | 30 epochs (hold-out) | 76.58 |
| "u-net" with batch-normalisation | 30 epochs (hold-out) | 82.19 |
Background and References
1. Marrone, S., Piantadosi, G., Fusco, R., Petrillo, A., Sansone, M., & Sansone, C. (2016, December). Breast Segmentation using Fuzzy C-Means and anatomical priors in DCE-MRI. In Pattern Recognition (ICPR), 2016 23rd International Conference on (pp. 1472-1477). IEEE.
2. Marrone, S., Piantadosi, G., Fusco, R., Petrillo, A., Sansone, M., & Sansone, C. (2013, September). Automatic lesion detection in breast DCE-MRI. In International Conference on Image Analysis and Processing (pp. 359-368). Springer Berlin Heidelberg.
3. Piantadosi, G., Fusco, R., Petrillo, A., Sansone, M., & Sansone, C. (2015). LBP-TOP for volume lesion classification in breast DCE-MRI. In 18th International Conference on Image Analysis and Processing, ICIAP 2015. Springer Verlag.
4. Long, J., Shelhamer, E., & Darrell, T. (2015). Fully convolutional networks for semantic segmentation. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (pp. 3431-3440).
5. Ronneberger, O., Fischer, P., & Brox, T. (2015). U-net: Convolutional networks for biomedical image segmentation. arXiv preprint arXiv:1505.04597.